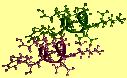 Using BTCL and TCL Commands To
Manipulate the Geometry of Two Helices, Instructions
Using BTCL and TCL Commands To
Manipulate the Geometry of Two Helices, Instructions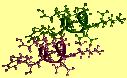 Using BTCL and TCL Commands To
Manipulate the Geometry of Two Helices, Instructions
Using BTCL and TCL Commands To
Manipulate the Geometry of Two Helices, Instructions
A few hints to understanding the script are:
% discovery helix
% ls -ltOutput files of the run should include helix.out, helix.cor, helix.arc, and helix.tbl
Search for these output lines (links are included to the hypertext version of the helix.out file):
axis1 = line {{11.0926 17.2649 3.27548}} {{-0.821539 0.123481 0.55662}} axis2 = line {{5.90624 11.5912 -3.12062}} {{-0.821539 0.123481 0.55662}} line distance = point 10.0 point distance = point 10.0The lines for axis1 and axis2 show the centroid, (the first { } in each line) and the direction vector (the second { } in each line) of the axes. Note that axis1 and axis2 have the same direction, indicating that they are parallel after execution of the fixorientation procedure. The line distance is the distance between two lines, and the point distance is the distance between two centroids. These two distances are the same, indicating that the centroids are lined up perpendicular to both axes at the predefined distance apart.
You can display the structures in the helix.cor file with the Insight program. Start up the Insight program from the directory in which you ran the Discover program by entering insightII at the UNIX prompt. Read in the helix.cor file with the File/Import command, and use Geometrics/Vector command of the DeCipher module to create the axes of the two helices by finding the least-squares fit line of the backbone atoms subset specified by molecule_name:*:ca,n,c.
(Before doing this, you might also want to display the molecules in the helix.car file to show that the helices in the original file were randomly oriented.)
Use the Data/Sort command to sort the data, by setting the following parameters:
 Main
access page
Main
access page  Advanced-Use (BTCL) access
Advanced-Use (BTCL) access
 BTCL - Tutorial Access.
BTCL - Tutorial Access. Copyright Biosym/MSI