This file is a container file supplied for convenient use. It combines together a number of files which contain the standard geometry for many types of chemical groups. The values in these files were entered over 15 years ago and, while complete, are not completely current. These are the default values used in TNT refinement because of their completeness.
The individual components of tntgeo_v010.dat are:
- protgeo.dat
This library contains the geometry restraints for the twenty amino acids, the peptide bond (PEPTIDE), the C-terminus (CTERM), and the disulfide bond (SS). The amino acids are identified by the usual three letter codes. To maintain compatibility with other programs the amino acid proline can be identified either as PRO or CPR. (Often CPR is used to identify a proline with a cis peptide bond. TNT treats an amino acid the same regardless of the conformation of the peptide so this distinction is not necessary.)
- bcorrel.dat
This file contains the B correlation restraints for the groups defined in protgeo.dat. When you wish to restrain the thermal parameters of your model because of the lack of high resolution data you should INCLUDE this file in addition to protgeo.dat. I have yet to create a corresponding file for the other geometry libraries, due to the lack of high resolution models from which the proper restraints can be derived.
- nuclgeo.dat
This file contains the geometry restraints for DNA and RNA. The five bases are identified by three letter codes. The sugar-phosphate backbone is linked with SUGPHOS in RNA and DSUGPHOS in DNA. The first base (at the 5' end) must be linked to the second via a 5'END or D5'END linkage. At the 3' end the final base must be linked to an empty residue via a 3'END or d3'END linkage. The atom names used in this library are those of the IUPAC, not those of the PDB. The atoms in the sugar rings are identified with primes ('), not stars (*).
- cofactor_geo.dat
This library contains the geometry restraints for ATP, ADP, AMP, CMP (i.e. cyclic AMP), dUMP, NAD, and NDP (i.e. NADPH). The atom naming convention is that of the PDB. The atoms in the ribose sugars are identified with stars (*). The PDB has chosen very strange names for the atoms in these groups, please read their documentation when attempting to use this library.
- othergeo.dat
This library contains geometry restraints for a number of small compounds. These include
- BENZENE Benzene
- BME Reduced
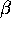 -Mercaptoethanol. Use SS to link two
BME residues or a CYS residue to a BME.
-Mercaptoethanol. Use SS to link two
BME residues or a CYS residue to a BME. - CBZ The N-terminal blocking group carbobenzoxy. Use CBZLINK to link a CBZ residue to the N-terminal of a peptide.
- DELTA Lyscine residue with all atoms past CD removed.
- DMSO Dimethyl Sulfoxide
- DTT Self-oxidized DTT
- DTT2 One half of a DTT. This residue type is used when a DTT residue sits on a crystallographic two-fold. Use the link DTTLINK to connect the DTT2 to itself.
- EPSILON Lyscine residue with all atoms past CE removed.
- GAMMA Lyscine residue with all atoms past CG removed.
- HEDS Two BME residues linked together. The atoms are named using the PDB convention.
- MSE Seleomethionine
- SO4 Sulfate (-2)
- chlorgeo.dat
This library contains the geometry definitions for bacteriochlorophyll-a.
This library contains the geometry definitions for carbomethoxy-heme (COHM).
A new search of the Cambridge Structure Database was performed by Engh and Huber (Engh, R.A., Huber, R. Acta Cryst (1991) A47, 392-400). The results of this survey have been used to construct a new geometry library for proteins. The restraints are consistent with the CSDX library in the XPLOR refinement package. However it is not consistent with the nucleic acid, and co-factor libraries described above. The result of mixing the two sources of restraint information is difficult to predict. If you have a simple protein this library can be used with no fear of problem.
Since the choice of restraint library will cause slight differences in the resulting model you should note the name of the library in your Protein Databank submission and in your paper.